Getting Started¶
System Requirements¶
mc3 is compatible with Python 3.6+, and has been tested to work on Unix/Linux and
OS X machines, with the following software:
- Numpy >= 1.19.5
- Scipy >= 1.9.3
- Matplotlib >= 3.3.4
mc3 may work with previous versions of these software;
however, we do not guarantee nor provide support for that.
Install¶
To install mc3 run the following command from the terminal:
pip install mc3
Or if you prefer conda:
conda install -c conda-forge mc3
Alternatively (e.g., for developers), clone the repository to your local machine with the following terminal commands:
git clone https://github.com/pcubillos/mc3
cd mc3
pip install -e .
mc3 provides MCMC and nested-sampling posterior sampling,
optimization and other lower-level statistical and plotting
routines. See the full docs in the api or through the Python
interpreter:
import mc3
# Bayesian posterior sampling:
help(mc3.sample)
# Optimization:
help(mc3.fit)
# Assorted stats:
help(mc3.stats)
# Plotting utilities:
help(mc3.plots)
Example 1: Interactive Run¶
The following example shows a basic MCMC run from the Python interpreter, for a quadratic-polynomial fit to a noisy dataset:
import numpy as np
import mc3
def quad(p, x):
"""
Quadratic polynomial function.
Parameters
p: Polynomial constant, linear, and quadratic coefficients.
x: Array of dependent variables where to evaluate the polynomial.
Returns
y: Polinomial evaluated at x: y = p0 + p1*x + p2*x^2
"""
y = p[0] + p[1]*x + p[2]*x**2.0
return y
# For the sake of example, create a noisy synthetic dataset, in a real
# scenario you would get your dataset from your data analysis pipeline:
np.random.seed(3)
x = np.linspace(0, 10, 100)
p_true = [3.0, -2.4, 0.5]
y = quad(p_true, x)
uncert = np.sqrt(np.abs(y))
data = y + np.random.normal(0, uncert)
# Initial guess for fitting parameters:
params = np.array([10.0, -2.0, 0.1])
pstep = np.array([0.03, 0.03, 0.05])
# Run the MCMC:
func = quad
output = mc3.sample(
data, uncert, func, params, indparams=[x],
pstep=pstep, sampler='snooker', nsamples=1e5, burnin=1000, ncpu=7,
)
That’s it. The code returns a dictionary with the MCMC results.
Among these, you can find the posterior sample
(posterior), the best-fitting values (bestp),
the lower and upper boundaries of the 68%-credible region (CRlo
and CRhi, with respect to bestp), the standard deviation of
the marginal posteriors (stdp), among other variables.
mc3 will also print out to screen a progress report every 10% of
the MCMC run, showing the time, number of times a parameter tried to
go beyond the boundaries, the current best-fitting values, and
lowest \(\chi^{2}\); for example:
::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::
Multi-core Markov-chain Monte Carlo (mc3).
Version 3.1.0.
Copyright (c) 2015-2023 Patricio Cubillos and collaborators.
mc3 is open-source software under the MIT license (see LICENSE).
::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::::
Yippee Ki Yay Monte Carlo!
Start MCMC chains (Wed Mar 29 17:52:45 2023)
[: ] 10.0% completed (Wed Mar 29 17:52:46 2023)
Out-of-bound Trials:
[0 0 0]
Best Parameters: (chisq=112.6196)
[ 3.12005211 -2.51498098 0.50946457]
Gelman-Rubin statistics for free parameters:
[1.05031159 1.04920662 1.05254562]
...
[::::::::::] 100.0% completed (Wed Mar 29 17:52:51 2023)
Out-of-bound Trials:
[0 0 0]
Best Parameters: (chisq=112.5923)
[ 3.07675149 -2.50001621 0.50890678]
Gelman-Rubin statistics for free parameters:
[1.00024775 1.00029444 1.00023952]
All parameters converged to within 1% of unity.
MCMC Summary:
-------------
Number of evaluated samples: 100002
Number of parallel chains: 7
Average iterations per chain: 14286
Burned-in iterations per chain: 1000
Thinning factor: 1
MCMC sample size (thinned, burned): 93002
Acceptance rate: 28.36%
Parameter name best fit median 1sigma_low 1sigma_hi S/N
--------------- ----------- ----------------------------------- ---------
Param 1 3.0768e+00 3.0761e+00 -3.7968e-01 3.8946e-01 7.9
Param 2 -2.5000e+00 -2.4981e+00 -2.2876e-01 2.1325e-01 11.2
Param 3 5.0891e-01 5.0868e-01 -2.6467e-02 2.7415e-02 18.7
Best-parameter's chi-squared: 112.5923
Best-parameter's -2*log(posterior): 112.5923
Bayesian Information Criterion: 126.4078
Reduced chi-squared: 1.1607
Standard deviation of residuals: 3.00577
For a detailed summary with all parameter posterior statistics see
mc3_statistics.txt
Output sampler files:
mc3_statistics.txt
At the end of the MCMC run, mc3 displays a summary of the MCMC
sample, best-fitting parameters, credible-region boundaries, posterior
mean and standard deviation, among other statistics.
Additionally, the user has the option to generate several plots of the MCMC
sample: the best-fitting model and data curves, parameter traces, and
marginal and pair-wise posteriors (these plots can also be generated
automatically with the MCMC run by setting plots=True).
The plots sub-package provides the plotting functions:
# And now, some post processing:
import mc3.plots as mp
import mc3.utils as mu
# Output dict contains the entire sample (posterior), need to remove burn-in:
posterior, zchain, zmask = mu.burn(output)
# Set parameter names:
pnames = ["constant", "linear", "quadratic"]
post = mp.Posterior(posterior, pnames)
# Plot pairwise posteriors:
fig_pairs = post.plot(savefile="quad_pairwise.png")
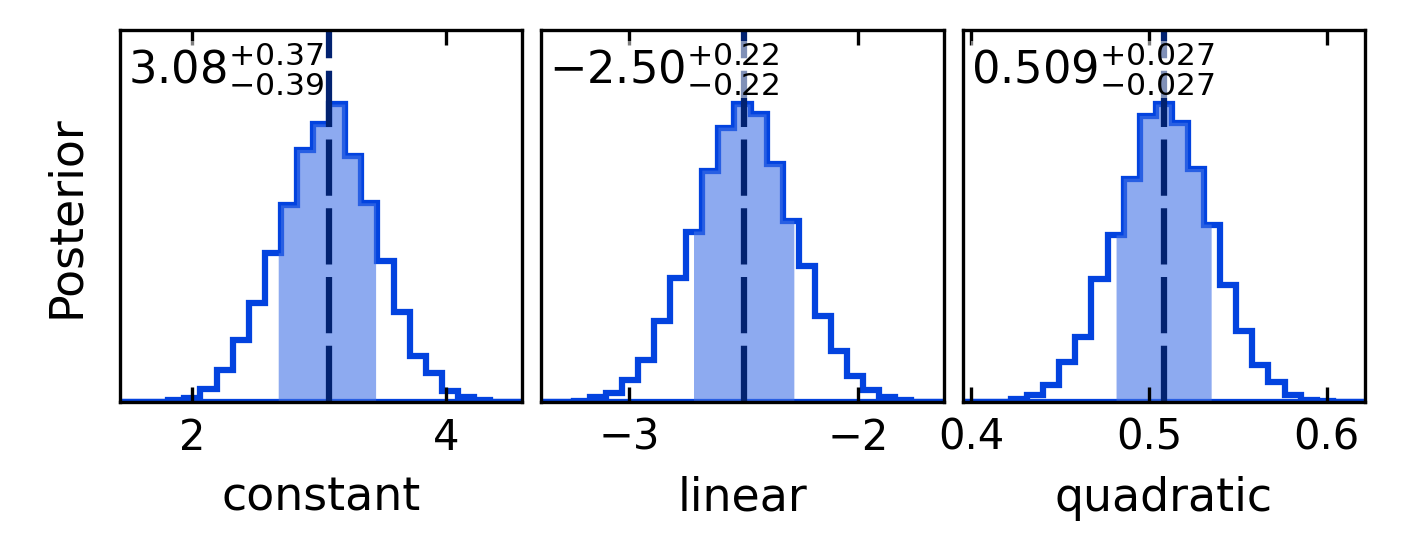
# Plot marginal posterior histograms:
fig_hist = post.plot_histogram(savefile="quad_hist.png")
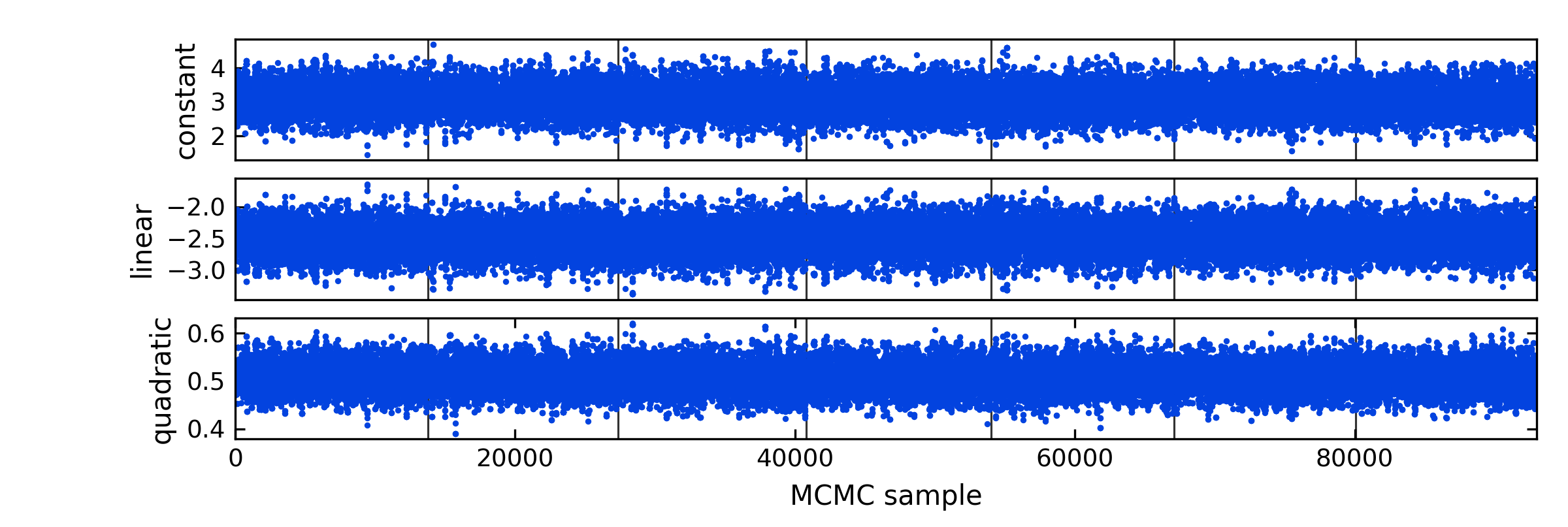
# Plot trace plot:
mp.trace(posterior, zchain, pnames=pnames, savefile="quad_trace.png")
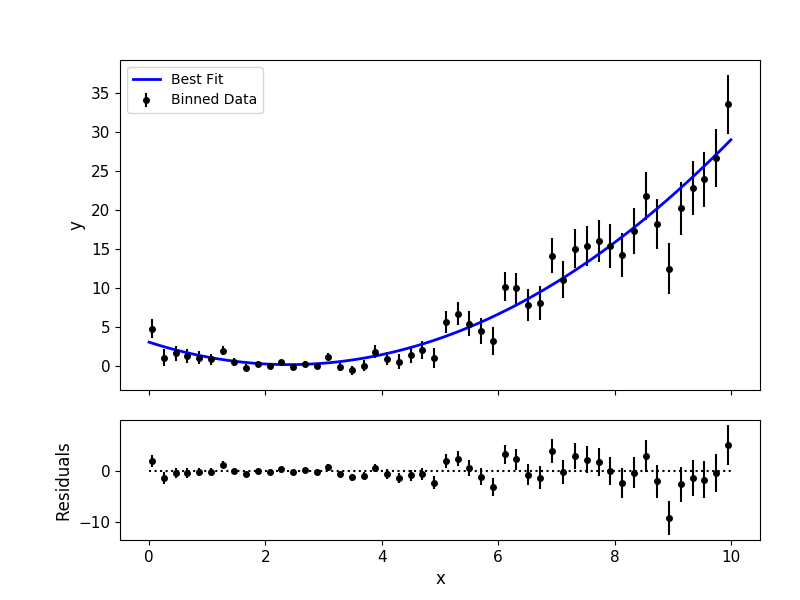
# Plot best-fitting model and binned data:
mp.modelfit(data, uncert, x, y, savefile="quad_bestfit.png")

Note
These plots can also be automatically generated along with the MCMC run (see Outputs).
Example 2: Shell Run¶
The following example shows a basic MCMC run from the terminal using a configuration file. First, create a Python file (’quadratic.py’) with the modeling function:
def quad(p, x):
y = p[0] + p[1]*x + p[2]*x**2.0
return y
Then, generate a data set and store into files, e.g., with the following Python script:
import numpy as np
import mc3
from quadratic import quad
# Create synthetic dataset:
x = np.linspace(0, 10, 1000) # Independent model variable
p0 = [3, -2.4, 0.5] # True-underlying model parameters
y = quad(p0, x) # Noiseless model
uncert = np.sqrt(np.abs(y)) # Data points uncertainty
error = np.random.normal(0, uncert) # Noise for the data
data = y + error # Noisy data set
# Store data set and other inputs:
mc3.utils.savebin([data, uncert], 'data.npz')
mc3.utils.savebin([x], 'indp.npz')
Now, create a configuration file with the mc3 setup (’MCMC.cfg’):
[MCMC]
data = data.npz
indparams = indp.npz
func = quad quadratic
params = 10.0 -2.0 0.1
pmin = -25.0 -10.0 -10.0
pmax = 30.0 10.0 10.0
pstep = 0.3 0.3 0.05
nsamples = 1e5
burnin = 1000
ncpu = 7
sampler = snooker
grtest = True
plots = True
savefile = output_demo.npz
Finally, call the mc3 entry point, providing the configuration file as
a command-line argument:
mc3 -c MCMC.cfg